When people talk about housing affordability, they are not always talking about the same thing. There are three distinct housing affordabilities:
- purchase affordability - how much it costs to buy a new or existing house at the time of the purchase,
- repayment affordability- how much it costs to service a mortgage on a house, and
- rental affordability- how much it costs to rent a house.
In this post I want to talk about the one that will be important over the next 18 months: repayment affordability.
As interest rates were lowered following the COVID pandemic, these were capitalised into higher housing prices. This is a long run trend, since the peak home loan rates in 1990. As lower and lower interest rates have allowed borrowers to borrow more money, this has contributed to higher and higher house purchase prices. [Note, this is not the only contributor to growing house prices; for example, long-run growth in two income households has also been important].

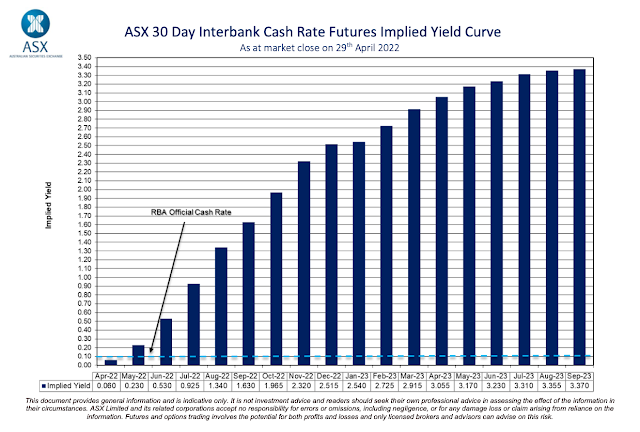
Higher inflation means the Reserve Bank of Australia (RBA) will lift interest rates to bring downward pressure on inflation. The ASX RBA rate tracker indicates there is a one-third probability that interest rates will increase on Tuesday this week. Rates will almost certainly rise by the first Tuesday in June. Over the next 18 months, the markets think that the RBA cash target rate will rise from the current target of 0.1 per cent to just over 3.3 per cent. That is a substantial increase.
What does this mean for variable rate home loans? The short answer is that they are likely to increase by a similar amount. By December they may be around 2.5 per cent higher, and by June next year they may be 3.2 per cent higher.
According to the ABS, the average new home loan for an owner occupier was \$595,873 in February 2022, down from \$620,315 in January 2022. According to the RBA, owner-occupier variable home loan rates are typically around 2.5 per cent, per year.
A 30 year, \$600,000 loan, with an interest rate of 2.5% has monthly repayments of \$2371. At a notional 5 per cent home loan borrowing rates in December 2022, the repayment increases to \$3221 per month. And at 5.7 per cent in September 2023, the repayment would be \$3482 per month.
Even for people whose loan is half that amount, they will experience a dramatic increase in household costs. A 25 year, \$300,000 loan, at a 2.5% interest rate has a monthly repayment of \$1346 per month. At 5% interest rate, the repayment becomes \$1754. At 5.7% the monthly repayment is \$1878.
If inflation breaks out as the market expects, and the RBA reacts as the market expects, repayment affordability will become the big concern of middle Australia over the next 18 months.
Ironically, when interest rates on home loans appreciate, house prices are likely to depreciate. This means that purchase affordability should improve over the next 18 months (albeit, not as much as some would hope as house prices are often what economists call sticky-downwards). Reduced house prices will leave some recent home-buyers with a difficult decision. Do they stay with an increased mortgage they are finding difficult to afford, or do they sell their new house and take a loss because house prices have fallen (what is good for the buyer is bad for the seller).